 
I'm choosing to use summarize_at instead of summarize and across because the specification of na.rm = TRUE as a separate argument to the across function is deprecated. Here's what my final function looks like after implementing their framework. Thanks to deschen for the solution using map2 from purrr to wrap the summarize call. I'm trying to avoid using for loops to keep this code compact and readable. Key R functions and packages The dplyr package v> 1.0.0 is required. before = Ozone) %>%Īny suggestions are greatly appreciated. This article describes how to compute summary statistics, such as mean, sd, quantiles, across multiple numeric columns. I'm using the built in R dataset "airquality" library(dplyr)ĭplyr::mutate(time = time_series. Here are some inputs and reproducible code to test with. # Set column names and delete extraneous columns created by the summarize functionĭplyr::summarize(dplyr::across(.cols = dplyr::all_of(variable_names), 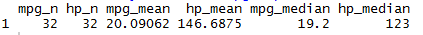
Here is a simple example of my current code: aggregate_func %ĭplyr::mutate(hour = lubridate::floor_date(time, "hour")) %>%ĭplyr::summarize_at(.vars = variable_names, I'm worried about the performance impact all these unnecessary computations will have. So far I have lived with this and just trimmed out the extra columns, but this function is going to be used on very large datasets (down to minute-by-minute interval data over multiple years) with a potentially large number of independent variables. For the example above, this means sum would be applied to all 4 columns, as would median, mean, and sd, resulting in 16 columns. When dplyr's summarize function is provided a list of variables and functions, it will apply every function to every column, squaring the total number of output columns. For example, if we have four independent variable columns, Ozone, Solar.R, Wind, and Temp, then the list of functions c(sum, median, mean, sd) would have Ozone aggregated by sum, Solar.R by median, Wind by mean, and Temp by standard deviation. The intention is that each function is only applied to the column it matches the order of. The number of functions should match the number of independent variable columns in the input dataframe and be in an order that corresponds to the order of columns to apply the functions over. 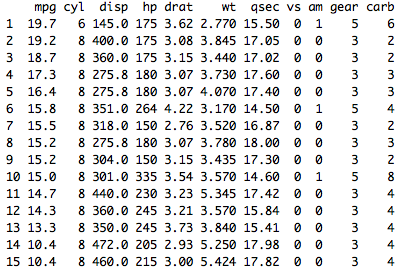
This is an argument to my function called variable_aggregation which is a vector of functions like c(sum, median, mean, sd). I want the user to be able to specify which functions are going to be used to aggregate data. For example, of those who are college graduates, how many are stem So far I have something like this. How do I go about calculating the proportion of a response for a certain subset of a data set. The input dataframe to my function has a time column and one or more independent variable columns. tidyverse dplyr xbechtel September 30, 2020, 3:16am 1 Hello I am very new to R. I am currently using dplyr and lubridate to accomplish this. For example, we would to apply n_distinct() to species, island, and sex, we would write across(c(species, island, sex), n_distinct) in the summarise parentheses.I am writing a function right now that will aggregate (roll up) data at short time intervals up to longer time intervals. n_distinct() in the example above, this external function is placed in the. When dplyr functions involve external functions that you’re applying to columns e.g. cols specifies the columns that you want the dplyr function to act on. It is used inside your favourite dplyr function and the syntax is across(.cols. Wouldn’t it be nice if we could just write which columns we want to apply n_distinct() to, and then specify n_distinct() once, rather than having to apply n_distinct to each column separately? Ordinarily, if we want to summarise a single column, such as species, by calculating the number of distinct entries (using n_distinct()) it contains, we would typically writeĭistinct_species distinct_island distinct_sex 
The new across() function turns all dplyr functions into “scoped” versions of themselves, which means you can specify multiple columns that your dplyr function will apply to. The first two columns, species and island, specify the species and island of the penguin, the next four specify numeric traits about the penguin, including the bill and flipper length, the bill depth and the body mass. There are 344 rows in the penguins dataset, one for each penguin, and 7 columns. # … with 334 more rows, and abbreviated variable names ¹flipper_length_mm, Species island bill_length_mm bill_depth_mm flipper_…¹ body_…² sex year 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
